Reliable assays for diagnosing COVID-19 disease are in strong demand in order to cope with the high number of subjects to be tested. Simon Mügge, PhD (Biochemistry) and European Patent Attorney, provides an introduction into the different approaches taken in diagnostic assays for COVID-19, demonstrating that the current pandemic promotes innovation and offers new opportunities to companies developing diagnostic assays and devices.
As of September 30, 2020, the WHO reports more than 33,5 million confirmed cases of COVID 19 and 1,004,421 deaths worldwide. In the absence of a safe and efficient vaccine, complying with hygiene rules, early detection and the quarantine of infected individuals are the only known preventive measures to stop further spreading of the disease. The following article provides basic insights into in vitro diagnostics for COVID-19.
COVID-19 virus can either be directly detected or indirectly by searching for antibodies against the virus
The novel beta coronavirus SARS-CoV-2, the causative pathogen of COVID-19, is a large RNA virus having a high similarity to the virus causing SARS (SARS-CoV) and MERS (MERS-CoV). SARS-CoV-2 has four main structural proteins: the spike (S) protein, nucleocapsid (N) protein, envelope (E) protein, and membrane protein (M). The N protein binds to the RNA genome and forms a capsid around the enclosed RNA. The other three proteins are essential for pathogenesis and viral replication, as well as genome maintenance.
Roughly, assays for the diagnosis of SARS-CoV-2 can be distinguished in assays (1) aiming for directly detecting the virus itself in a sample of the tested subject, and (2) assays aiming for detection of antibodies in a sample of the tested subject, indirectly indicating that the subject’s immune system has been challenged with SARS-CoV-2.
Assays aiming for directly detecting the virus itself have the advantage that they can detect presence of SARS-CoV-2 already during the early stage of disease, when the subject’s immune system has not yet formed antibodies against the virus. On the other hand, it is reported that in severe cases of COVID-19, the virus RNA does not disappear as fast from the body as in non-severe cases.
The RT-PCR technique amplifies virus RNA, if present
The most widely used technique in this respect aims at the detection of the virus’ RNA using a method called Reverse-transcription polymerase chain reaction (RT-PCR).
In a first step, the RNA of the virus is reverse transcribed into complementary DNA (cDNA) using an enzyme called reverse transcriptase. The term ‘reverse’ expresses that this process runs reverse to the normal process in a cell, where DNA is used as a template to produce a RNA copy, which is then exported from the cell’s nucleus (a messenger RNA, mRNA) and translated into protein.
The cDNA obtained in the RT-step is then mixed with short DNA-strands, so called primers. These primers are designed to align and bind to sequences which are unique for the new SARS-CoV-2 virus. Typically, primers are selected to detect different regions of the virus RNA, such as regions coding for the N protein and regions for the E protein, in order to increase the specificity of the assay. If the cDNA contains a sequence, which is complementary to the sequence of a pair of primers, another enzyme (polymerase) can multiply the DNA sequence between said two primers, thereby providing a detectable DNA fragment of a defined and expected size.
Each replication step requires applying a cycle of heating and cooling to the reaction mixture, and in case DNA is replicated, its number is doubled at the end of the temperature cycle. The number of temperature cycles required for detection of said DNA fragment thus allows an quantitative estimation of the amount of virus (virus load) present in a sample. Therefore, said method can be rendered a quantitative RT-PCR (qRT-PCR or RT-qPCR).
Eventually, the replicated sample is analysed by way of an end-point control for the presence of the replicated DNA fragment. Nowadays, specific dyes can be used in the PCR amplification step, which detect the increase of double-stranded DNA, i.e. the increasing number of the replicated DNA fragment, in real time.
Another variation of this assay exists, in which additional DNA probes (so called Taq-Man probes) are added. These probes are also designed to be complementary to the replicated DNA fragment. Upon binding to the DNA sequence being replicated it releases a fluorescence dye, which is subsequently detected by a fluorescence reader. While the RT-PCR method is specific and sensitive, it takes 2.5 to 3 hours, and requires sample collection, transport and handling by trained personnel. At the same time, companies such as the Finnish based company Mobidiag developed a closed cartridge system, which simplifies the handling and further speeds up the assay.
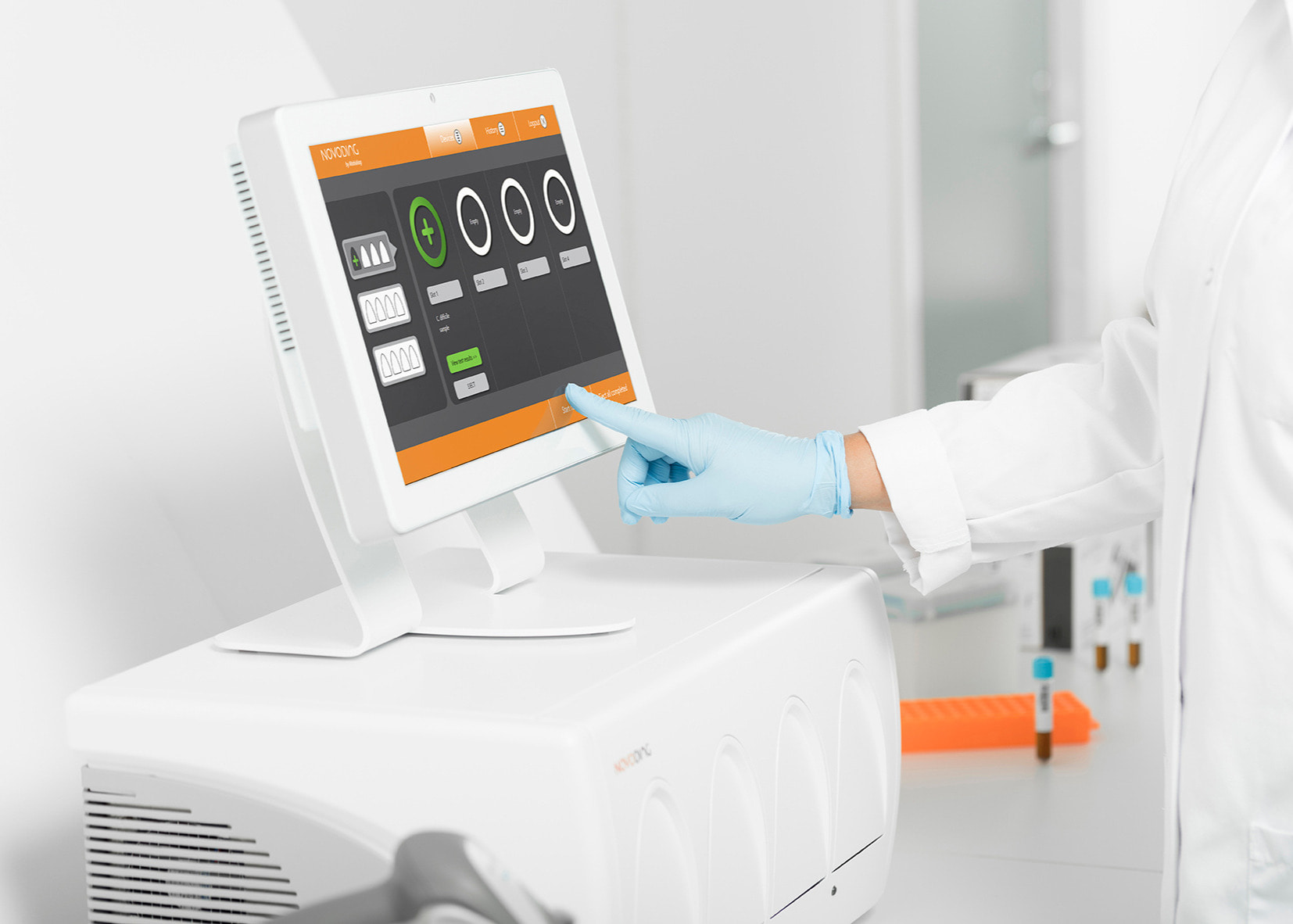

A closed cartridge test system Novodiag® by Mobidiag.
Newly developed methods avoid the time-consuming temperature cycling step of RT-PCR
Additional methods of nucleic acid replication have been developed, which avoid the time-consuming heating and cooling steps. Instead, after the required reverse transcription step, these assays are designed to amplify DNA with high efficiency at a single temperature.
One recently approved method is the loop-mediated isothermal amplification method (LAMP), which is able to provide results as fast as in 15-60 minutes and does not require expensive reagents or instruments. However, since no temperature cycles are used, LAMP provides a qualitative result, reporting on the presence or absence of the virus only. Detection is made using fluorescence dye binding to the amplified DNA, like in real-time RT-PCR.
Other LAMP detection techniques use small probes which are labelled with a dye and a small molecule called biotin. Following incorporation of these probes into the replicated DNA fragment, these molecules can be ‘fished’ out of the sample solution using the biotin-binding molecule streptavidin. This eventually allows that replicated DNA can be made visible to the naked eye as a red line on a test strip.
Another LAMP assay uses quenching probes (Qprobes). Such QProbes are small DNA molecules carrying a fluorescence dye. Upon hybridization with its target nucleotide sequence, the fluorescence is quenched by electron transfer between the fluorophore and a guanine residue in the target, reducing the fluorescence signal emitted by the probe.
A recently developed new method uses the CRISPR associated enzyme Cas13 for rapid and portable sensing of SARS-CoV-2 RNA. Cas13 is an enzyme which can be programmed to selectively bind to a target sequence, here the amplified RNA. When activated, Cas13 cleaves a probe comprising both a fluorophore and a quencher, thereby releasing the fluorophore, which in turn can be detected. Still, further optimisation of this technique is required to enable large-scale evaluation.
Still another assay widely used in the detection of coronavirus is the microarray assay. In this assay, SARS-CoV-2 RNA is first reverse transcribed into cDNA labelled with specific probes. In a second step, these labelled cDNAs are incubated with oligonucleotides fixed on a well of a plate (the microarray) such that they bind to the microarray. Following several washing steps to remove unbound cDNA, the virus RNA can be detected by detection of the specific probes.
ELISA tests detecting virus protein allows high throughput testing
Direct detection of SARS-CoV-2 is not limited to detection of viral RNA. Rather, several immunoassays (assays using antibodies for detection) are used for detecting SARS-CoV-2 protein in patient samples such as respiratory tract samples, serum, plasma or whole blood. The virus proteins function as antigens, which are only detected when the virus is replicating. Hence, these tests can be used to identify acute or early infection.
Among these, the most common setup used is an ELISA (enzyme linked immunosorbent assay). In this setup, an antibody specific for the virus protein is first attached to a plate (coating). A sample is incubated with the plate, allowing the virus protein to be captured by the antibody. Following several washing steps to remove unbound material, a second antibody labelled with an enzyme for detection is incubated with the plate. In case virus protein was captured, said detection antibody will also bind to the virus protein, and the labelled enzyme drives in a detection step a colour reaction, and the colour formed can be measured.
The chemiluminescence assay uses a similar set-up, but detection is made via chemiluminescence. Automated sample handling allows high throughput testing with detection rates equal or higher than those of nucleic acid amplification techniques.
Assays targeting for antibodies against SARS-CoV-2 do not allow detection of early infection
As mentioned in the beginning, a second group of assays aims for detection of antibodies in a blood sample of the tested subject, thereby indirectly indicating that the subject’s immune system has been challenged with SARS-CoV-2. While the handling of blood samples is easier as compared to the above-mentioned methods, these antibody detecting assays are limited to the avail that IgM antibodies first start to develop from 4-10 days after infection, and IgG antibodies from 10-20 days post infection. On the other hand, the amount of IgM and IgG antibodies also provides a pointer towards the time of initial infection and can be used to monitor the course of infection.
One very common experimental set-up is again the ELISA format. However, instead of a capture antibody, virus protein is first coated to the plate, such that antibodies specifically recognizing the virus protein are captured and later-on detected using chemiluminescence or an enzyme driven colour-formation.
An even simpler setup is the lateral flow immunochromatographic assay (LFIA), a format which is also commonly used in pregnancy tests. The sample to be analysed diffuses through a strip, in which detection antibodies are embedded, resulting in the formation of an antibody-antigen complex. The complex is then captured by a line of antibodies, which are fixed in their position. If virus protein is present, a coloured line becomes visible.
Advantageously, these assays can be carried out as simply as with a blood drop obtained from a prick in the finger. It does not require expensive equipment or specially trained personnel, and therefore qualifies for testing of large number of patients. However, not all tests are CE Marking approved and widely available. Further information on immunoassays targeting COVID-19 is provided, for example, by Foundation for Innovative New Diagnostics (FIND).
Non-specific tests can aid in diagnosing COVID-19
Finally, it should also be mentioned that – in addition to SARS-CoV-2 specific tests – also non-specific routine-tests are used in COVID-19 diagnosis, which focus on the change of parameters or combination of parameters, which are symptomatic for COVID-19.
Suitable tests may be directed towards changes in levels of cytokines (cellular transmitters evidencing inflammatory states), changes in blood coagulation, changes in the composition and numbers of white blood cells, or enzymes indicating cellular damage in organs. A further discussion of these assays would, however, go beyond any boundaries of this article.

False-positives in COVID-19 diagnostics
Accuracy is a measure how well an assay correctly identifies or excludes a condition. Often this requires setting a threshold within a ‘grey’ area. As a result, values obtained from an infected person may in some cases lie below said threshold such that the person is incorrectly diagnosed as being healthy (a false negative result). Likewise, in other cases values obtained from a non-infected person may lie above the threshold such that the person is incorrectly diagnosed as being infected (a false-positive result).
In case it is of importance to detect all truly infected persons to inhibit spreading of a disease (i.e. to reduce the number of false-negatives), a conservative approach can be taken by setting the threshold, above which tests are reported positive, rather low. Consequently, the number of false-positive tested healthy persons will increase.
Social media recently spread the argument that the increasing number of positive tested COVID-19 cases is merely due to a false-positive rate of tested persons as high as 15%. According to the argument, said 15% of all tests would account for the increase in reported positive tested cases. However, said false positive rate should be correctly applied to the number of positive tested cases only. An increase in the number of positive tested cases thus means that 85% of said positive tested cases are truly infected. Accordingly, even with a percentage of 15% false positive, an increasing number of positive tested cases still truly reports an increase of infections in the tested population.
Taking into account that a conservative approach significantly lowers the number of false-negative tests, and that in case of doubt patients can be re-tested, a certain false-positive rate may be considered acceptable.
Kolster supports companies in the development of innovative diagnostics with a team of experts in the field of life science and experts with deep knowledge in the field of diagnostic and medical devices. Kolster assists in taking innovation-driven opportunities in Finland and Europe, and – in cooperation with its trusted partners – world-wide. In case of interest, we are happy to evaluate with you how Kolster’s service can be of value to your business.
Get to know Kolster's biotech and chemistry team.
Simon Mügge
Partner, German & European Patent Attorney, European Trademark and Design Attorney
simon.mugge@kolster.com
+358 40 920 8814
